Follicle Atresia
Vertebrate follicle atresia represents the death of an organized follicle at any time during development beginning with the primordial stage through the ovulatory stage. Atresia is generally considered a normal physiological process by which surplus or non-viable growing follicles are rapidly reabsorbed by a non-inflammatory process, though it can also be abruptly induced by environmental or physiological perturbations. It is significant to note, however, that among vertebrate species the overall proportion of ovarian follicles lost by atresia, as well as the time during follicle development at which atresia can occur, differs according to reproductive strategy. In particular, those animals that produce thousands to perhaps millions of mature gametes within a reproductive cycle (spawners; e.g. fish) show a very low overall incidence of atresia, and most of the oogonia produced during the lifespan of the female are ovulated as mature oocytes. By comparison, those groups that produce a considerably more limited number of offspring (many reptiles, birds, mammals) demonstrate a proportionately higher incidence of atresia. Furthermore, within a normal reproductive cycle a major difference between reptiles and birds compared to mammals is that in the former species, atresia occurs prior to selection into the preovulatory hierarchy, whereas in mammals atresia occurs following selection of the dominant follicle(s).
Avian Follicle Atresia Mediated via Apoptosis
In birds, the loss of follicles by atresia can occur any time during the active breeding season, at the termination of breeding (following the onset of photorefractoriness or at the start of incubation), and during seasonal molt, and the overall rate of atresia has been observed to increase with advancing age. Under optimal breeding conditions atresia rarely occurs in preovulatory follicles, but can be induced at this final stage of development by inappropriate environmental conditions (e.g., during forced molt induced by food deprivation and/or reduction in photoperiod) and a decline in circulating gonadotropins. By contrast, follicle atresia does occur in primary and prehierarchal follicles, though the exact incidence and cause(s) have yet to be established. Thus, it has been proposed that a transition from atresia-susceptibility to atresia-resistance is initiated coincident with follicle selection into the preovulatory hierarchy (Johnson, 2003). In the domestic hen, the comparatively higher incidence of follicle atresia in prehierarchal follicles, in vivo, correlates with susceptibility of the granulosa cell layer to undergo apoptosis as observed, in vitro (Johnson, Bridgham, Witty and Tilly, 1996). In light of this observation, and because apoptosis in granulosa cells represents the earliest event described during the onset of atresia, most studies of the ovary have focused primarily on this cell type.
The following represents a simplified model of extrinsic and intrinsic mechanisms and pathways that promote apoptosis in hen granulosa cells (for details, see Johnson, 2003 and Johnson and Woods, 2007).
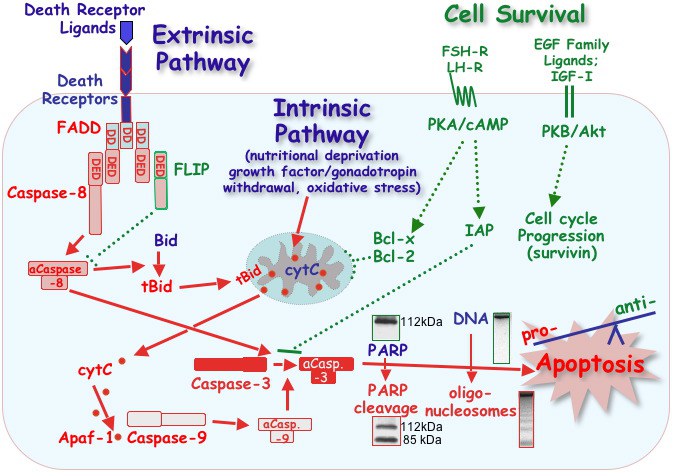
Simplified model of pro- and anti-apoptotic pathways in hen granulosa cells. Apoptotic cell death can be initiated by either an intrinsic or extrinsic pathway, both of which converge at the activation of caspase-3. Activation of this executioner enzyme is irreversible, and results in the cleavage of numerous structural and functional proteins (e.g., PARP) and the internucleosomal cleavage of genomic DNA. Cell death pathways are opposed by anti-apoptotic proteins that protect mitochondrial membrane integrity (Bcl-x, Bcl-2), block caspase activity (IAPs) or prevent signaling via death receptors (the adaptor protein, FLIP). Expression of these anti-apoptotic proteins is regulated by gonadotropins (FSH and LH) and locally produced growth factors (IGF-I and EGF family ligands) via cell survival signaling pathways. The absence of sufficient anti-apoptotic protein expression (dotted green lines) is proposed to tip the balance in favor of activating pro-apoptotic pathways (solid red lines). Abbreviations: aCasp.-3, -7, -8, activated caspases; Apaf-1, Apoptosis Protease Activating Factor-1; Bid and tBid, intact or truncated BH3 Interacting Domain death agonist; cytC, cytochrome C; DD, death domain; DED, death effector domain; EGF, epidermal growth factor; FADD, Fas-Associated Death Domain; FLIP, Flice-like inhibitory protein; FSH, follicle-stimulating hormone; Inhibitor of Apoptosis Protein, IAP; IGF-I, insulin-like growth factor I; LH, luteinizing hormone; PARP, Poly-(ADP-ribose)-polymerase; PKA/cAMP, protein kinase A/cyclic AMP signaling pathway; PKB/Akt, protein kinase B/Akt signaling pathway. Each of the proteins and cell signaling pathways depicted has been characterized in hen granulosa cells, in vitro. For a more complete description with citations, see Bridgham et al., 2003; Johnson, 2003 and Johnson and Bridgham, 2002.
Death Receptors (DRs) Mediating Apoptosis via the Extrinsic Pathway
Extracellular factors, in particular those from the tumor necrosis factor (TNF) family of ligands, bind to cognate death domain-containing members of the tumor necrosis factor receptor family (death receptors) which then indirectly initiate the caspase cascade. Intracellular death domains of activated death receptors recruit and bind cytoplasmic adaptor proteins via homophilic interactions, while adaptor proteins (e.g., Fas associated via death domain, FADD), in turn, interact with initiator caspases (e.g., caspase 8) through a death effector domain (DED). The processing of caspase 8 subsequently activates downstream caspases directly as well as initiates the mitochondrial signaling pathway described above, thus amplifying the initial signal and assuring eventual cell death. Our laboratory has taken a comparative approach to understanding this process by utilizing the chicken (Gallus gallus) ovary model system. Perhaps not surprisingly, results thus far demonstrate that there exists a striking conservation of intracellular survival and cell death-mediating pathways between mammalian and avian species.
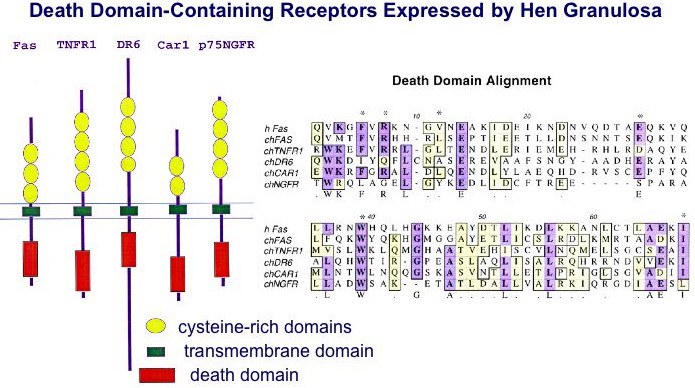
Common features of all death receptors include multiple cysteine-rich repeats within the extracellular ligand-binding domain, a type I (single pass) transmembrane domain, and a cytoplasmic death domain required for intracellular signaling. The CAR1 receptor is proposed to represent the Gallus ortholog to the mammalian TRAIL receptor, DR5 (Bridgham and Johnson, 2002). For additional information, see: Bridgham and Johnson, 2001; Bridgham, Bobe et al., 2001 and Bridgham, Wilder et al., 2003.
Phylogenetic Relationships for Death Receptors Among Vertebrates
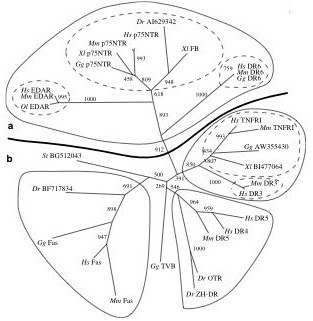
Phylogeny of the death receptor gene family based upon death domain homology. This tree was constructed using the neighbor-joining algorithm of the program NEIGHBOR in the PHYLIP (phylogeny inference package) program. Distances were calculated using the Jones-Taylor-Thornton similarity matrix. Bootstrap support (out of 1000) is indicated. A full maximum likelihood tree (estimated using PROML from the PHYLIP package) estimated the same topology. In addition to the phylogeny shown, we searched for all death domains found in other non-receptor proteins using the tBLASTn search. Including these death domains, found in both vertebrate and invertebrate species, in our analysis did not alter the topology or increase the resolution of the phylogeny for the receptor proteins, nor did they provide evidence of a single ancestral death domain (data not shown). A and B denote different functional categories (see text for explanation). Hs=Homo sapiens, Mm=Mus musculus, Gg=Gallus gallus, Xl=Xenopus laevis, St=Silurana tropicalis, Dr=Danio rerio, Ol=Oryzias latipes. For additional details, see Bridgham, Wilder, Hollocher and Johnson, 2003.
Expression of Granulosa Cell Proteins Responsible for Mediating Cell Death via the Extrinsic Pathway
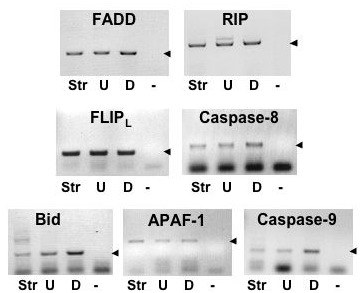
Amplification of cDNAs corresponding to TNFRSF adaptor proteins (FADD and RIP), extrinsic pathway proteins (anti-apoptotic protein, FLIPL, and caspase-8) and intrinsic pathway pro-apoptotic proteins (Bid, APAF-1 and caspase-9) expressed in undifferentiated (U) granulosa cells from 6-8 mm follicles and differentiated (D) granulosa from preovulatory follicles (arrow heads). Ovarian stromal tissue (Str) is included as a reference tissue. - , negative control (no template). Johnson et al., 2007.
TRAIL Signaling in Granulosa Cells
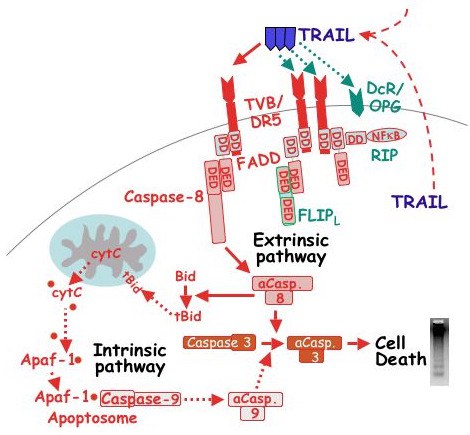
Proposed pathway for TRAIL signaling in hen granulosa cells. Solid arrows depict activation of the extrinsic pathway via the putative Gallus ortholog to DR5 (TVB; Bridgham & Johnson, 2002). Red dotted arrows depict the predicted activation of the intrinsic pathway based upon the demonstrated expression of APAF-1 and caspase-9 mRNA in granulosa cells from both prehierarchal and preovulatory follicles. Dotted green arrows predict potential pathways by which TRAIL-induced apoptotsis is prevented, possibly via FLIPL or decoy receptors (DcR), including osteoprotegerin (OPG; Bridgham & Johnson, 2003). In addition, the potential exists for TRAIL to activate the receptor adaptor protein, RIP. Because TRAIL mRNA is normally expressed within ovarian stromal, thecal and granulosa tissues, it is predicted that TRAIL can act on granulosa cells in a paracrine and/or autocrine fashion (dashed lines). aCasp., activated caspase; DD, death domain; DED, death effector domain; tBid, truncated Bid. For additional details, see Johnson et al., 2007).
Anti-Apoptotic Proteins Mediating Resistance of Granulosa Cells to Apoptosis
Expression and attenuation of apoptosis by the Bcl-2 family member, Bcl-xLong

Left panels. Representative sections showing the distribution of Bcl-xLong protein in hen preovulatory (A) and prehierarchal (B) follicles by immunocytochemical analysis. Panels C and D represent controls following incubation with preimmune serum. Note the prominent presence of Bcl-xLong protein (arrow heads) in cytoplasm of granulosa (Gr) from preovulatory, but not prehierarchal, follicles. By comparison, there is some degree of Bcl-xLong staining in prehierarchal follicle theca interna (TI) and theca externa (TE). Y, yolk. Magnification x 600.
Right panels. Over-expression of Gallus g. Bcl-xLong in Chinese hamster ovary (CHO) cells attenuates caspase-9-like activity and the progression of apoptosis in staurosporine-treated cells. CHO cells were transfected with Gallus Bcl-xLong (cBcl-xL) and clonally selected for stable expression (top panel). Cultured cells were treated with 0 to 200 nM staurosporine for 18 h, then collected for evaluation of oligonucleosome formation (middle panel) or assayed for caspase-9-like activity using the fluorogenic substrate, Ac-LEHD-AFC. * denotes a significant attenuation of in vitro caspase activity compared to the Control cells. For additional details, see Johnson, Bridgham and Jensen, 1999 and Johnson and Bridgham, 2002.
Gallus Survivin-142 and its splice-variants
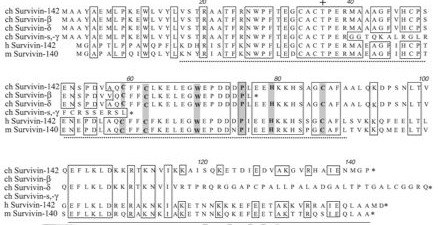
Deduced amino acid sequence for chicken (ch) Survivin-142 and its alternatively spliced forms (Survivin-beta [accession #AJ394392], -delta, -gamma, and -short [s]) aligned to human (h) Survivin-142 (accession #U75285) and murine (m) Survivin-140 (accession #AB013819). Numbering corresponds to the ch Survivin, the dotted underlined region indicates the predicted BIR domain, while the solid underline represents the putative a-helical coiled-coil motif. Boxes show amino acids which are completely conserved among the ch, h and m Survivin sequences, while shaded residues represent the various residues reported to be critical for BIR domain function (16,29). +, Thr phosphorylation site found to be phosphorylated by p34cdc2-cyclin B1 and reported to be essential for maintaining cell viability at cell division (O'Connor et al., Proc. Nat. Acad. Sci. 97:13103, 2000); *, stop codon.
Gallus Survivin-142 as a cell cycle-related and antiapoptotic protein
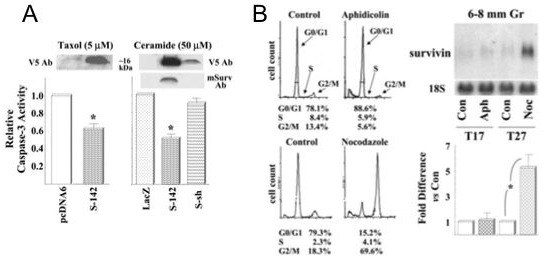
A. Top panels: Transfection of primary cultured hen granulosa cells with full-length Survivin (S-142) or Survivin short (S-sh). Representative Western blots of protein from transfected hen granulosa cells using a V5 epitope antiserum (V5 Ab) which recognizes both the ch Survivin-142 and -short fusion proteins, or an anti-mouse Survivin-140 serum (mSurv Ab). Note that levels of Survivin protein are not readily detected in control-transfected cells and that, as predicted, the anti-mouse Survivin serum does not recognize the transfected ch Survivin-short protein. Bottom panels: Transient transfection of hen granulosa cells with chicken Survivin-142 attenuates caspase-3 activity following treatment with 5 mM Taxol for 20 h (left panel) or 50 mM ceramide for 6 h (right panel). By comparison, chicken Survivin-short (S-sh) which lacks the complete BIR domain, fails to attenuate ceramide-induced caspase-3 activity. *, P<0.05 versus the corresponding control (either the pcDNA6/V5-His vector alone or vector containing LacZ). B. Differential expression of survivin mRNA relative to stage of cell cycle. Granulosa cells from 6-8 mm follicles were cultured with TGFalpha for 17 h (T17) in the absence or presence of aphidicolin (Aph, 30 mM; to block at G0/G1), or for 27 h (T27) in the absence or presence of nocodazole (Noc, 1.3 mM; to block at G2/M). Cells were subsequently collected for cell cycle analysis by flow cytometry, and the percentage of cells at each stage found in a representative experiment are indicated (left panels). In addition, cells were processed for quantitation of survivin mRNA (right panels). *: P<0.05. For additional details, see Johnson, Langer and Bridgham, 2002.

